 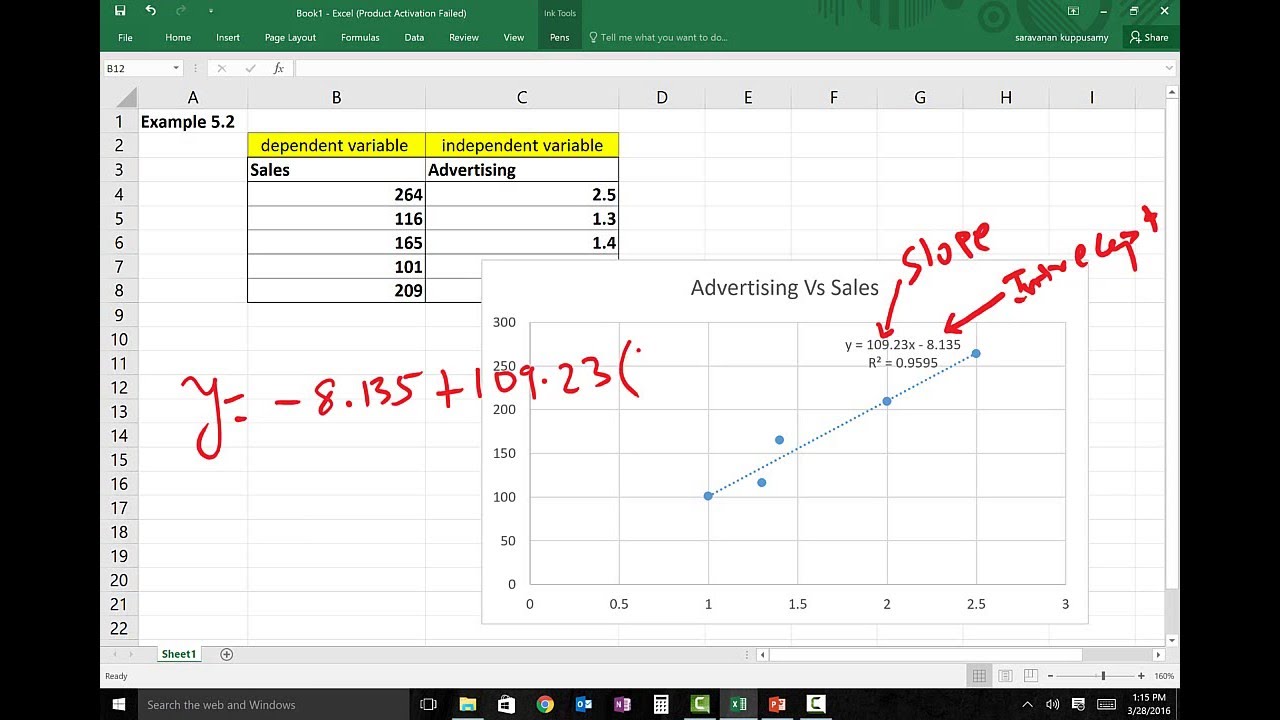
This is an overall significance test assessing whether the group of independent Independent variables does not reliably predict the dependent variable. 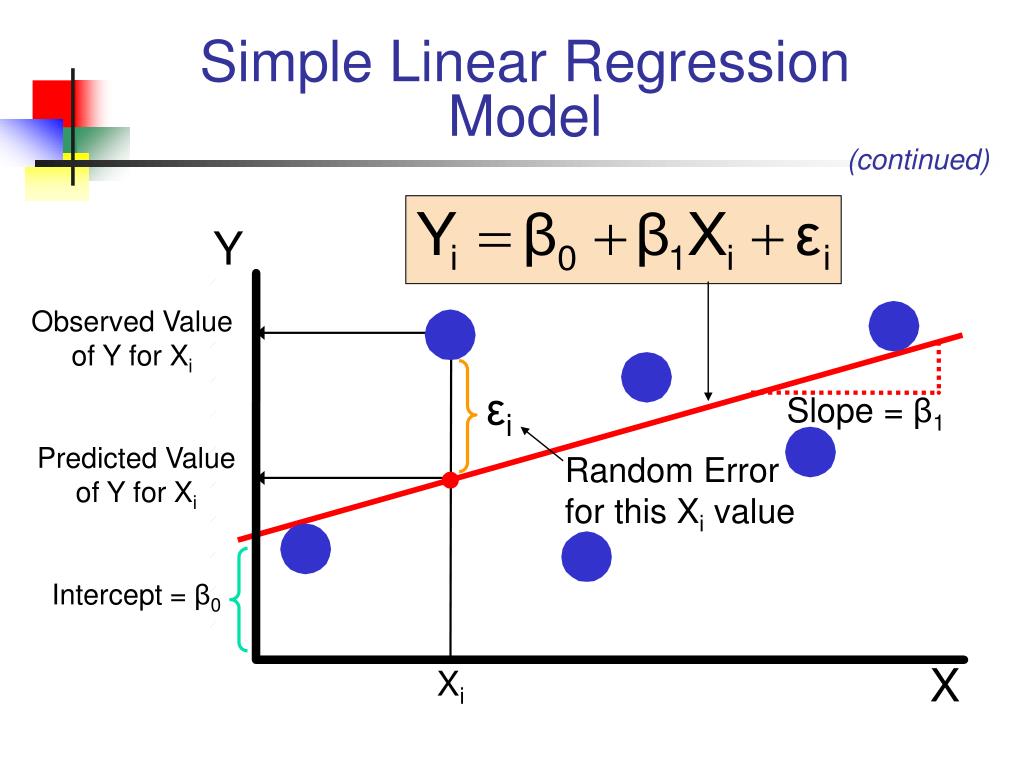
Statistically significant relationship with the dependent variable, or that the group of If the p-value were greater thanĠ.05, you would say that the group of independent variables does not show a Reliably predict science (the dependent variable). That the group of variables math, female, socst and read can be used to Independent variables reliably predict the dependent variable”.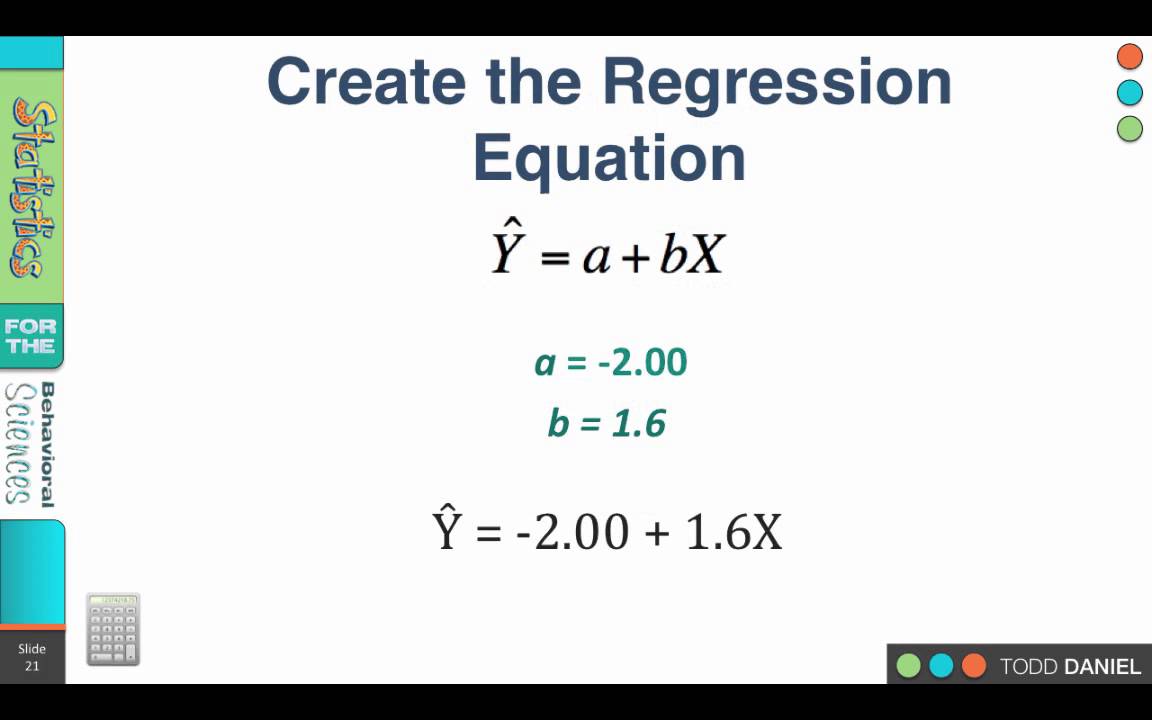 
The p-value is compared to yourĪlpha level (typically 0.05) and, if smaller, you can conclude “Yes, the Reliably predict the dependent variable?”. These values are used to answer the question “Do the independent variables The p-value associated with this F value is very small (0.0000). Model science = math female socst read / clb ĭependent Variable: science science score Analysis of Variance Sum of Mean Quit tells SAS that not to expect another proc reg immediately. Statement is included because proc reg is an interactive procedure, and The clb option after the slash on the model statement to get theĩ5% confidence limits of the parameter estimates. With the dependent variable (in this case, science) on the left of theĮquals sign, and the independent variables on the right-hand side. The model statement, we specify the regression model that we want to run, Tells SAS where to find the SAS data set to be used in the analysis. In the code below, the data = option on the proc reg statement The variable female is a dichotomous variable coded 1 if the student was Scores on various tests, including science, math, reading and social studies ( socst). These data ( hsb2demo) were collected on 200 high schools students and are I think the use of the CMPTMODEL statement would enable fitting all of the data, but I will have to learn more about it before I post anything.This page shows an example regression analysis with footnotes explaining the Ods output parameterestimates=parms additionalestimates=addl Pred = c0*trt1*exp(k0*trt1*(ATPTN-43)) + (c0 + d1)*trt2*exp((k0 + d2)*trt2*(ATPTN-43)) Įstimate 'removal coefficient for trt1' k0 Įstimate 'removal coefficient for trt2' k0 + d2 Įstimate 'half-life for trt1' -log(2)/k0 Įstimate 'half-life for trt 2' -log(2)/(k0 + d2) Proc nlmixed data=have2(where=(atptn>40)) maxiter=5000 However, in many pharmacokinetic studies, this is the standard approach, and it gives a good estimate of the removal coefficient and the resulting half-life in the pool. However, that estimate of t1/2 will be biased by not having an accurate initial estimate of the maximum concentration and when it occurs. That would be easiest by removing time points less than 43 from the analysis and rescaling the time axis. One thing that might help would be to estimate the removal coefficient and estimate t1/2 from that. Neither of those models fit the linearized (log transform) approach you have. There are two logical models that fit this - a time independent two-pool model with sampling from the second pool, so that there is an entry rate constant, or a time-dependent model with sampling from a single pool. It appears that at time=0 the response value is at or near zero, then increases to about time = 43, followed by a decrease. So I started to work on this with NLMIXED when I noticed something critical - these data do not describe a single pool decaying process of the type c = c0 * exp (-k * time). Output out= t_half mean= Mean_T_HALF min= Min_T_HALF max= Max_T_HALF Proc means data= reg_mod(where= (Variable='ATPTN')) If Variable = 'ATPTN' then T_HALF = log10(0.5)/Estimate Input USUBJID ACTARMN ATPTN AVAL 1đđ5Ĕ9 1đĒ9ė9 Label USUBJID = "Unique Subject Identifier" *H0: Humoral immune response in Group 1 as assessed by PRNT half-life will be similar to Group 2*/ Can log10 transformations be used instead?Ģ) If so, then the model equation would be log10(AVAL)=log10(AVAL0)+k*t, where AVAL is the PRNT results, AVAL0 is the PRNT results at baseline, k is the reaction rate constant, and t is the time (days).ģ) Then the slope would be k? So half-life can be calculated using t_1/2=log10(0.5)/k.Ĥ) Is the following code an accurate way to calculate half-life and p-value? I'm not confident in my methodology, and would appreciate some verification/feedback.ġ) Most of what I've seen in calculating half-life use natural log-transformations. I want to calculate half-life by fitting a linear regression model to log transformed PRNT titers, then obtain p-values using Mann-Whitney U Test. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
